Purpose
As a result of this lab, you will be able to...
Overview
The sense of “taste” results from complex interactions among the signals that our brain receives from our nose (“smell”) and the signals that our brain receives from the taste receptors located in taste buds in our tongue. Taste receptors are proteins that recognize substances in food.
PTC (phenylthiocarbamide) is a chemical that is considered “bitter” in taste. This chemical is not found naturally in foods. Why would our genome need to have a gene that would determine our ability to taste this chemical? Scientists are not completely sure. PTC has a molecular structure that is similar to chemicals found in poisonous plants. One thought is that this gene played a part in the taste receptors for these other poisonous chemicals. There seems to be a correlation between PTC tasting and the dislike of broccoli and related plants. (Broccoli is not poisonous, despite what you told your parents when you were little.)
Arthur Fox discovered PTC tasting ability in the 1930’s, when he synthesized phenylthiocarbamide (PTC) in his lab. While doing so, the PTC dust made a lab assistant comment that he could taste bitterness in the air. However, Fox could not taste the dust in the air. Why could one person taste it and another couldn’t? We now know it is because of their genes. Albert Blakeslee continued this research through taste testing and found that the inability to taste PTC is a recessive trait. For example, non-tasters would have the genotype tt, and strong tasters TT.
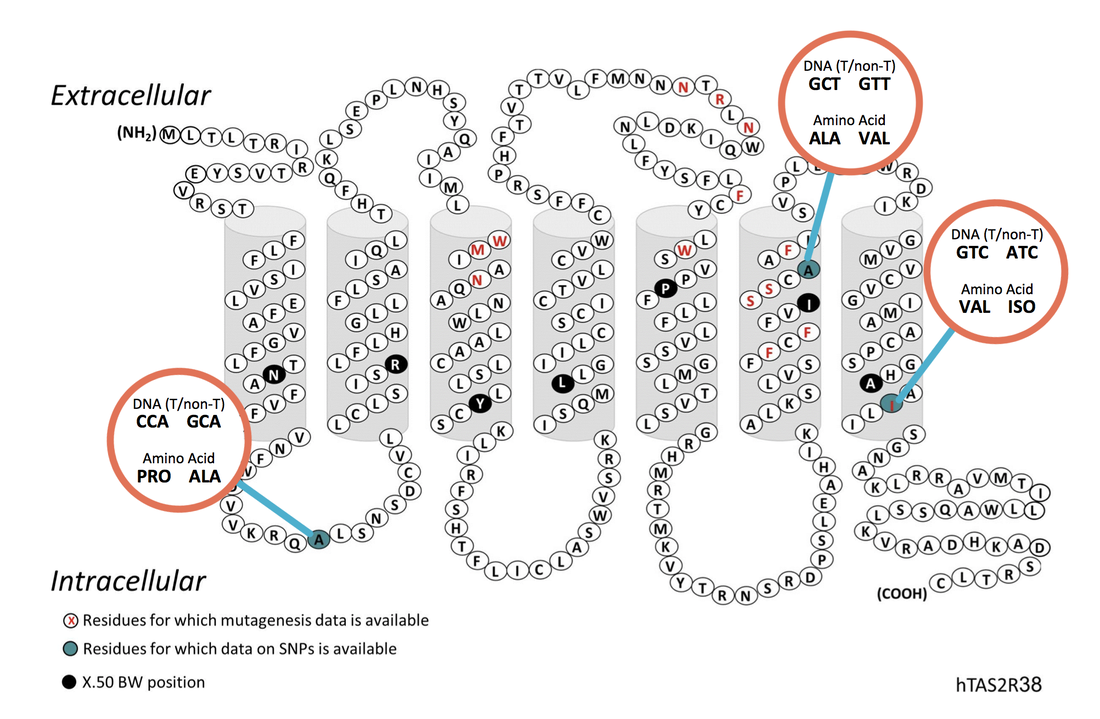
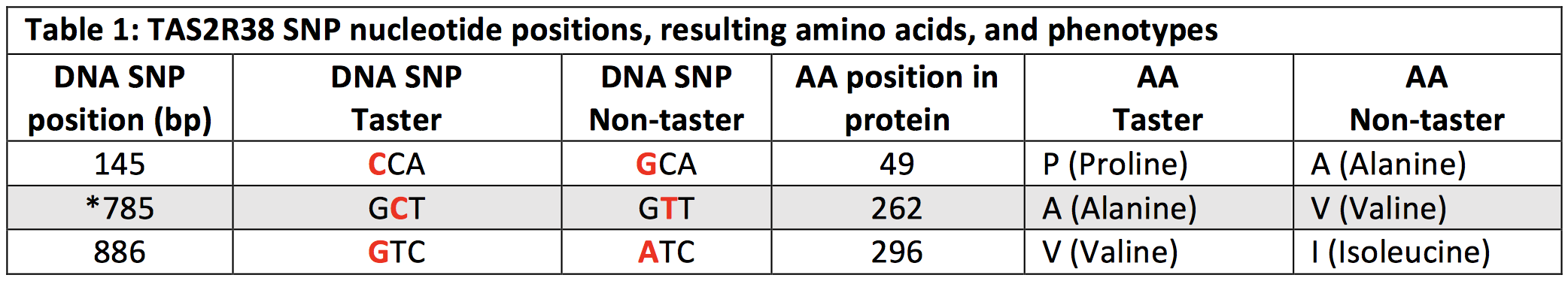
There are about 30 genes that encode bitter taste receptors in mammals. In 2003, scientists determined the sequence of the gene associated with PTC bitter tasting: taste receptor 2 member 38, or TAS2R38. This gene is 1,143 nucleotides long and has 3 locations where variations in the sequence correlate with the PTC tasting and non-tasting. Each of these changes is called a “SNP” (pronounced “snip”). A SNP is a single nucleotide polymorphism, or change in a single nucleotide in the sequence of DNA. We will use one of the SNPs (the one from position 785 in Table 1) to predict if you are a PTC “taster” or “non-taster.” A restriction enzyme can cut your DNA at position 785 if you have a taster allele but not if you have a non-taster allele!
DNA and Protein Sequences
The common non-taster allele has a G at nucleotide position 145 (G145), T at position 785 (T785) and A at position 886 (A886). This non-taster allele produces a protein with alanine, valine and isoleucine at these sites and therefore is also referred to as the AVI allele. The common taster allele has C145, C785, and G886, produces a protein with proline, alanine, and valine at these sites, and is referred to as the PAV allele.
As a result of this lab, you will be able to...
- Understand how a change in the sequence of a gene can result in a change in the function of the protein encoded by that gene, and consequently result in a changed phenotype.
- Use important molecular biology tools and techniques (DNA isolation, PCR, restriction enzyme digest of DNA,gel electrophoresis, and micropipetting) to determine your own genotype at the TAS2R38 locus.
Overview
The sense of “taste” results from complex interactions among the signals that our brain receives from our nose (“smell”) and the signals that our brain receives from the taste receptors located in taste buds in our tongue. Taste receptors are proteins that recognize substances in food.
PTC (phenylthiocarbamide) is a chemical that is considered “bitter” in taste. This chemical is not found naturally in foods. Why would our genome need to have a gene that would determine our ability to taste this chemical? Scientists are not completely sure. PTC has a molecular structure that is similar to chemicals found in poisonous plants. One thought is that this gene played a part in the taste receptors for these other poisonous chemicals. There seems to be a correlation between PTC tasting and the dislike of broccoli and related plants. (Broccoli is not poisonous, despite what you told your parents when you were little.)
Arthur Fox discovered PTC tasting ability in the 1930’s, when he synthesized phenylthiocarbamide (PTC) in his lab. While doing so, the PTC dust made a lab assistant comment that he could taste bitterness in the air. However, Fox could not taste the dust in the air. Why could one person taste it and another couldn’t? We now know it is because of their genes. Albert Blakeslee continued this research through taste testing and found that the inability to taste PTC is a recessive trait. For example, non-tasters would have the genotype tt, and strong tasters TT.
There are about 30 genes that encode bitter taste receptors in mammals. In 2003, scientists determined the sequence of the gene associated with PTC bitter tasting: taste receptor 2 member 38, or TAS2R38. This gene is 1,143 nucleotides long and has 3 locations where variations in the sequence correlate with the PTC tasting and non-tasting. Each of these changes is called a “SNP” (pronounced “snip”). A SNP is a single nucleotide polymorphism, or change in a single nucleotide in the sequence of DNA. We will use one of the SNPs (the one from position 785 in Table 1) to predict if you are a PTC “taster” or “non-taster.” A restriction enzyme can cut your DNA at position 785 if you have a taster allele but not if you have a non-taster allele!
DNA and Protein Sequences
The common non-taster allele has a G at nucleotide position 145 (G145), T at position 785 (T785) and A at position 886 (A886). This non-taster allele produces a protein with alanine, valine and isoleucine at these sites and therefore is also referred to as the AVI allele. The common taster allele has C145, C785, and G886, produces a protein with proline, alanine, and valine at these sites, and is referred to as the PAV allele.
Image Credit: Ayana Wiener; Marina Shudler; Anat Levit; Masha Y. Niv. BitterDB: a database of bitter compounds. Nucleic Acids Res 2012, 40
(Database issue):D413-419.
(Database issue):D413-419.
Polymerase Chain Reaction (PCR)
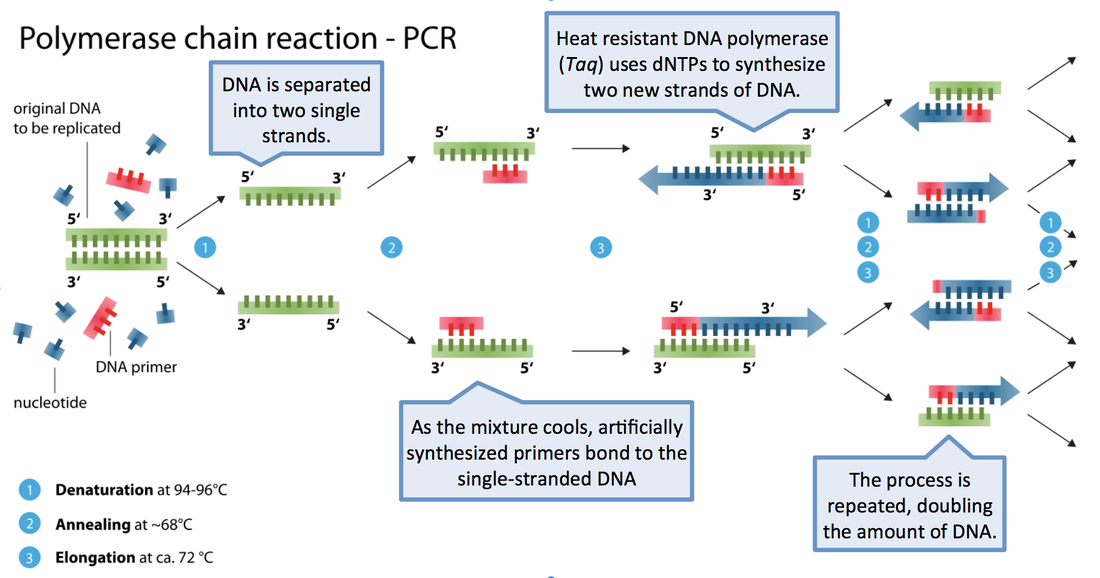
PCR is a quick, easy way to make many copies of a region of DNA from a small number of starting copies in a few hours. This technique is one of the most widely used techniques in molecular biology.
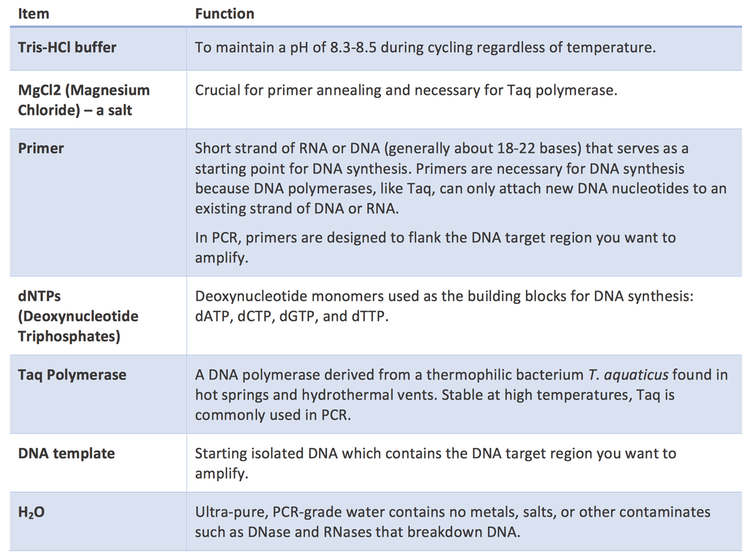
Components of a PCR reaction:
PCR is a quick, easy way to make many copies of a region of DNA from a small number of starting copies in a few hours. This technique is one of the most widely used techniques in molecular biology.
Components of a PCR reaction:
- Template DNA: this is the DNA that contains a region you are interested in amplifying.
- Nucleotides (dNTPS): these are the building blocks of DNA, which allow for synthesis of new DNA strands
during the PCR cycles. The name “dNTP” refers to the mixture of the four bases that make up DNA: deoxy-
ATP, dGTP, dTTP, and dCTP. - Primers: these are short pieces of DNA that are specific to the region of DNA that you would like to amplify.
The two primers in a PCR reaction must be complimentary to the beginning and end of the region you wish to amplify. It’s important that the primers are complimentary to opposite strands to allow for amplification of the DNA between the primers. - DNA polymerase: the workhorse of PCR. One of the most common polymerases used in PCR is “Taq.” Taq gets its name from the bacterium Thermus aquaticus, which lives in hot springs and can tolerate high temperatures. This enzyme is responsible for elongating the primers by adding complimentary nucleotides along the template strand of DNA, synthesizing the new strand of DNA.
- Buffer: provides the salts and is necessary to create the optimal working conditions for the DNA polymerase
- Magnesium Chloride (MgCl2): a cofactor required for the activity of DNA polymerase. Sometimes MgCl2 is
included in the buffer.
Steps of a PCR reaction:
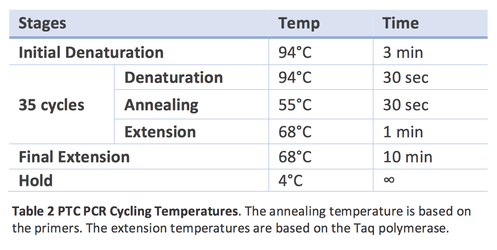
1. Denaturation, 94° C: Heating the DNA at high temperature breaks the hydrogen bonds between the two strands of DNA to make two single stranded molecules of DNA. This allows the DNA bases to be accessible for the primers to find their complimentary sequence in the next step.
2. Annealing, ~55-65° C: Cooling the single stranded DNA allows the primers to bind to their complimentary sequence. Choosing an appropriate annealing temperature can be one of the trickiest parts of PCR. Too low and primers can stick to non-specific sequences; too high and they may not bind at all. The optimal temperature can be approximated based on primer length and composition using online tools.
3. Extension, 72° C: This is the step when the DNA polymerase goes to work, adding nucleotides to the primers to synthesize a new strand of DNA.
From one to a million: PCR is an exponential process. Each cycle doubles the copy number of the region being amplified. After 20 cycles, you will have over a million copies of your single starting copy.
1. Denaturation, 94° C: Heating the DNA at high temperature breaks the hydrogen bonds between the two strands of DNA to make two single stranded molecules of DNA. This allows the DNA bases to be accessible for the primers to find their complimentary sequence in the next step.
2. Annealing, ~55-65° C: Cooling the single stranded DNA allows the primers to bind to their complimentary sequence. Choosing an appropriate annealing temperature can be one of the trickiest parts of PCR. Too low and primers can stick to non-specific sequences; too high and they may not bind at all. The optimal temperature can be approximated based on primer length and composition using online tools.
3. Extension, 72° C: This is the step when the DNA polymerase goes to work, adding nucleotides to the primers to synthesize a new strand of DNA.
From one to a million: PCR is an exponential process. Each cycle doubles the copy number of the region being amplified. After 20 cycles, you will have over a million copies of your single starting copy.
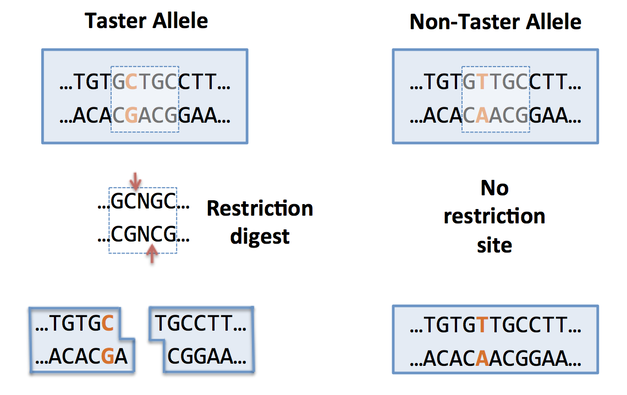
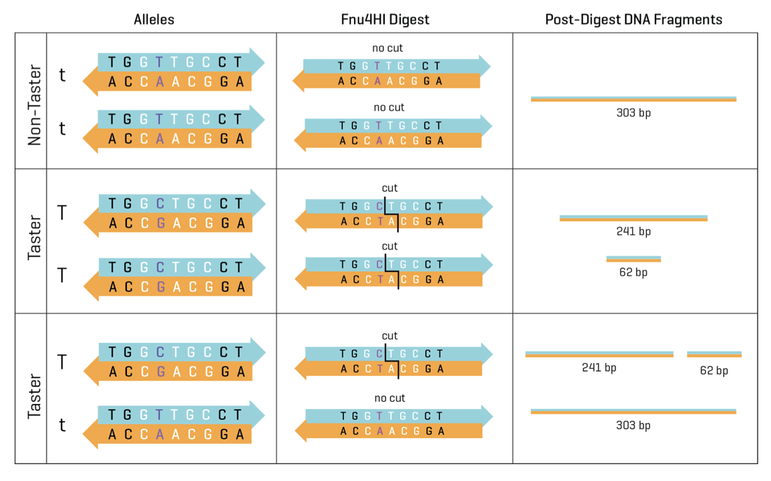
You will isolate DNA from your cheek cells and use PCR to copy a small region of the TAS2R38 gene using polymerase chain reaction (PCR). We will then use a restriction enzyme called Fnu4HI to perform a restriction digest on this PCR product. A restriction digest is the process of cutting a strand of DNA using an enzyme that recognizes a specific sequence (the “restriction site. Fnu4HI recognizes and cuts at GCNGC (“N” can be any nitrogen base). If your PCR product contains GCNGC, the enzyme will cut the PCR product into two pieces. This result will indicate that your gene has the code for the ability to taste PTC. If your gene doesn’t contain GCNGC, the enzyme will not cut, and you are homozygous recessive (tt) for this gene. You will compare your PCR product (303 base pairs) to your restriction digested PCR product using agarose gel electrophoresis.
Remember, you have 2 copies of the TAS2R38 gene: one from your mother and one from your father. Your PCR product might have some copies with the cutting site for the enzyme and some without. This would indicate that you are heterozygous with a genotype of Tt. Since “T” is dominant, you are a “taster.” However, your reaction to the PTC paper and to bitter substances will probably not be as strong as to someone who has is homozygous dominant. If all of the DNA copies cut into two pieces, you are homozygous dominant (TT) and are also a “taster.”
There is a chance you might not see the smallest fragment (62 bp) of our PCR products since it may run off the gel or be too faint to see. Using your ladder, which has DNA fragments of known lengths, will help you determine the size fragments you have. Detection of your genotype will mainly be from the other two larger fragments (303 for non- tasters, 241 for homozygous taster, or both fragments for heterozygous individuals).
Remember, you have 2 copies of the TAS2R38 gene: one from your mother and one from your father. Your PCR product might have some copies with the cutting site for the enzyme and some without. This would indicate that you are heterozygous with a genotype of Tt. Since “T” is dominant, you are a “taster.” However, your reaction to the PTC paper and to bitter substances will probably not be as strong as to someone who has is homozygous dominant. If all of the DNA copies cut into two pieces, you are homozygous dominant (TT) and are also a “taster.”
There is a chance you might not see the smallest fragment (62 bp) of our PCR products since it may run off the gel or be too faint to see. Using your ladder, which has DNA fragments of known lengths, will help you determine the size fragments you have. Detection of your genotype will mainly be from the other two larger fragments (303 for non- tasters, 241 for homozygous taster, or both fragments for heterozygous individuals).
Check your understanding:
1. Which trait, tasting or non-tasting is dominant?
2. Imagine you start with 1 copy of the gene, how many copies of the gene will you have after running 10 cycles of PCR?
3. Why don't we use human DNA polymerase in PCR if we are making copies of a human gene?
4. After doing the restriction digest we will run the DNA through gel electrophoresis. Draw what bands you would expect to see on the gel if the DNA came from someone that was a heterozygote.
1. Which trait, tasting or non-tasting is dominant?
2. Imagine you start with 1 copy of the gene, how many copies of the gene will you have after running 10 cycles of PCR?
3. Why don't we use human DNA polymerase in PCR if we are making copies of a human gene?
4. After doing the restriction digest we will run the DNA through gel electrophoresis. Draw what bands you would expect to see on the gel if the DNA came from someone that was a heterozygote.
/Volumes/group-1/*SEP Files/*Uber SEP = Index/*Curriculum Updates/2017 Curriculum Update/PCR/Tech Background 171023.docx 10/23/17
Procedure
Step 1: DNA Isolation from cheek cells
To isolate DNA from your cheek cells. The PTC taste receptors are encoded by the gene TAS2R38 located on chromosome 7.
Step 2: Polymerase Chain Reaction
To amplify (make many copies of) a particular portion of the TASR38 gene that contains a single nucleotide difference associated with differences in tasting PTC.
PCR fragment length: 303 base pairs (bp)
Step 3: Restriction Enzyme Digest
If the single nucleotide difference associated to PTC tasting is present in the post-PCR DNA, the restriction enzyme Fnu4HI will recognize the sequence and cut. If the post-PCR DNA has the non- tasting allele, Fnu4HI will not recognize the sequence and no cuts will occur.
Step 4: Gel Electrophoresis and visualization of results
To see if the enzyme cut the amplified DNA pieces or not. This will allow us to determine the genotype at that SNP.
After enzyme digest:
To isolate DNA from your cheek cells. The PTC taste receptors are encoded by the gene TAS2R38 located on chromosome 7.
Step 2: Polymerase Chain Reaction
To amplify (make many copies of) a particular portion of the TASR38 gene that contains a single nucleotide difference associated with differences in tasting PTC.
PCR fragment length: 303 base pairs (bp)
Step 3: Restriction Enzyme Digest
If the single nucleotide difference associated to PTC tasting is present in the post-PCR DNA, the restriction enzyme Fnu4HI will recognize the sequence and cut. If the post-PCR DNA has the non- tasting allele, Fnu4HI will not recognize the sequence and no cuts will occur.
Step 4: Gel Electrophoresis and visualization of results
To see if the enzyme cut the amplified DNA pieces or not. This will allow us to determine the genotype at that SNP.
After enzyme digest:
- Nontaster (tt): 303 bp band
- Taster (TT): 241 bp bands (there is also a 62 bp band that you may not see)
- Heterozygote (Tt): 303 bp, 241 bp, (and possibly 62 bp)
Step 1: Isolating DNA from Cheek Cells
Purpose: Isolate pure DNA that can be used for PCR from your cheek cells. Use a high heat and Chelex protocol to remove contaminates and deactivate enzymes that might breakdown DNA or inhibit PCR.
Purpose: Isolate pure DNA that can be used for PCR from your cheek cells. Use a high heat and Chelex protocol to remove contaminates and deactivate enzymes that might breakdown DNA or inhibit PCR.
- Label two microtubes and a cup for saline solution with your group number and initials.
- Pour about 5ml of saline into your cup (5 ml is about a pinky width high in our small cups)
- Pour saline solution in your mouth and vigorously rinse your cheeks for 1 minute.
- Expel the saline solution back into the cup.
- Carefully, over a garbage bin or sink, pour your spit and saline solution into one of your microtubes until it is almost completely full. If available use a sterile transfer pipet instead of pouring. To reduce the spread of flu or other pathogens, do not handle other people’s samples. Keep the remaining sample in the cup in case you don’t get a pellet after centrifuging.
- Close the microtube tightly and put it into a high-speed centrifuge. Spin for 8 minutes at to collect the cheek cells at the bottom of the microtube. (Wait for the centrifuge to fill before spinning. We have two for the whole class.)
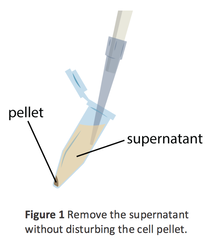
- After centrifuging, look for a cell pellet at the bottom of the tube. Carefully pour the liquid above the pellet (supernatant) into a waste container (if you have a pellet then your saliva container becomes your waste container) and keep the cell pellet in the tube. Repeat step 5 again if there is no pellet and remove the supernatant - pour waste down the drain.
- To inhibit DNA damage by DNAses and other enzymes, add 250 μl of 10% Chelex using a P1000 micropipet. Gently agitate the bottle of Chelex before pipetting as the beads will settle quickly to the bottom. The P1000 micropipet is with the Chelex.
- Use a Vortexer to resuspend the pellet in the Chelex solution.
- Once fully resuspended, close the tube lid tightly. To prevent the lid from popping open, use a microtube cap lock or heavy non-plastic material.
- Place the tube in a 99°C heat block for 10 min to breakdown cells and degrade contaminating proteins. (Wait until you have 6-8 tubes ready to be heated. Place the brick over the tubes in the heat block to prevent the lids from popping open as the tubes are heated.)
- After 10 min, resuspend the solution one more time by flicking the tube with your fingers (or Vortex).
- Once resuspended, place the tube in a high-power centrifuge. Spin for 6 minutes in a high speed centrifuge. (Wait for the centrifuge to fill before spinning. We have two for the whole class.) This will pull the heavier cell debris to the bottom, while the lighter DNA will remain in solution.
- Without disturbing the Chelex beads at the bottom of your microtube, pipet 50 μl of the supernatant into your second tube. Do not transfer any of the Chelex beads that could disable polymerase during PCR. This tube now contains your extracted DNA.
- Prepare a cup of ice. You will need it for the Performing PCR step as well as this step.
- Place your tube of extracted DNA on ice until you are given the next set of instructions. You will either move on to the PCR step if there is time or your teacher will store the extracted DNA in the freezer until the next class.
- Clean up: Dispose of all saliva cups, your first microtube containing the leftovers from the DNA extraction, and tips. Put everything back in your kit as you found it. You will need the tips and the DNA extraction tube for the next step, nothing else.
Step 2: Performing PCR
Purpose: Isolate and amplify a fragment of the gene of interest, TAS2R38, from your genomic DNA using PCR.
Purpose: Isolate and amplify a fragment of the gene of interest, TAS2R38, from your genomic DNA using PCR.
- Keep all reagents and DNA on ice throughout the procedure.
- Locate your second kit. It has a P20 micropipette, small tube rack, and a back of tiny tubes.
- Label your tiny microtube (0.2 ml) with your period, group #, initials, and contents of the tube (example: p1.g5. HC PCR) on the top and sides. This is your PCR reaction tube.
- Take your container of ice (with extracted DNA), tips, and your P20 micropipette to the teacher desk by the door. You will find the reagents you need there.
- Add the PCR reagents to your labelled PCR microtube. Use a P20 pipet and a new tip for every reagent.
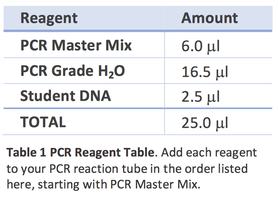
- Add 6 μl of PCR MasterMix into your PCR tube.
- Add 16.5 μl of PCR-grade water.
- Finally, add 2.5 μl of your DNA.
- Close the tube and mix by gently tapping the tube. Then, use the microcentrifuge to spin all the liquid to the bottom of your tube. (You have to put your small PCR tubes into a microtube so that it will not fall through the holes in the centrifuge. This has been set up for you but only one centrifuge is set up for the PCR tubes.)
- Return your PCR tube to ice. When the class is ready you will move your PCR tube to the PCR machine and load the machine according to the seating chart. Keep your groups' tubes together.
- While waiting, clean up your lab station. Return the tips to the first kit. Make sure everything is clean and back how you found it. The only thing you should have out is your ice cup with PCR tubes.
- While waiting, clean up your lab station. Return the tips to the first kit. Make sure everything is clean and back how you found it. The only thing you should have out is your ice cup with PCR tubes.
- You teacher will run the PCR machine according to the program, once all the tubes are in place.
Components of a PCR reaction – what’s in your tube?
Step 3: Restriction Enzyme Digest of PCR Product
Purpose: Use the restriction enzyme Fnu4HI to recognize and cut the DNA (PCR product) on a single nucleotide polymorphism (SNP) corresponding to the ability to taste PTC. The non-tasting allele (t) will not be recognized by Fnu4HI and will result in no cut.
Purpose: Use the restriction enzyme Fnu4HI to recognize and cut the DNA (PCR product) on a single nucleotide polymorphism (SNP) corresponding to the ability to taste PTC. The non-tasting allele (t) will not be recognized by Fnu4HI and will result in no cut.
|
Image Credit: Fred Hutchinson Science Education Partnership. http://libguides.fredhutch.org/ld.php?content_id=36813729
Step 4: Gel Electrophoresis Analysis of Restriction Digest
Purpose: Analyze your PCR and enzyme digest products by separating the DNA fragments according to size using gel electrophoresis.
Purpose: Analyze your PCR and enzyme digest products by separating the DNA fragments according to size using gel electrophoresis.
- Find another group of four that is ready to pour a gel.
- Pour a gel using preheated agarose. Make sure to put the comb in on the correct side. (DNA is negatively charged so the comb should be placed at the top of the gel, closest to the negative side.)
- Measure 30ml of agarose into the medicine cup (it will be almost full)
- Add 3μl of SYBR to the medicine cup with the liquid agarose. Stir until evenly mixed.
- Pour the agarose from the medicine cup into the casting tray (make sure the comb is at the top of the tray and that the 9 comb side is down). Fill both sides of the tray by pouring half of the agarose into each side.
- Remember: SYBR is light sensitive so after pouring the gel you need to cover the casting tray with aluminum foil.
- PAUSE HERE WHILE WE WAIT FOR THE RESTRICTION DIGEST TO COMPLETE.
- Put the gel box near the power supply or outlet before loading samples. Moving the box once samples are loaded may cause samples to wash out of the wells.
- Add enough 0.5X TAE buffer to cover the gel (approximately 150ml). The TAE buffer is already at your lab station.
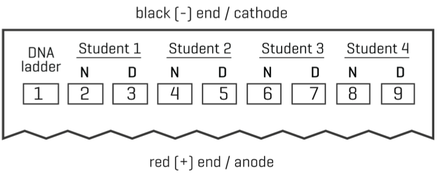
- You will be loading the DNA ladder, all group member’s digested DNA samples and each “no- digest” post-PCR product into the gel. Record in your lab notebook how your group will load each student’s digested and non-digested samples. (SEE THE DIAGRAM TO THE RIGHT). Note which samples belong to each group member.
- Obtain your digested (“D”) and no digest (“N”) tubes from the teacher.
- Pipet 2 μl of loading dye into your “D” tube and 2 μl of loading dye to your “N” tube to prepare your samples.
- Load your gel.
- Pipet 10 μl of the DNA ladder into the well designated by your Gel Diagram.
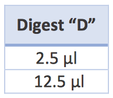
- Pipet 12 μl of your “D” sample into the well designated by your Gel Diagram.
- Pipet 12 μl of your “N” sample into the well designated by your Gel Diagram.
- When all lanes are loaded, connect your gel box to your power supply. Close the box and run the gel for 20-30 minutes check the box every 5 minutes to confirm that the system is working and that your dyes have not migrated off the end of the gel.
- After your DNA has migrated down 75% of the gel, turn off the gel box and unplug the power supply.
- There are dyes in the DNA ladder so you can see how far the DNA has travelled. Do not let it run off the end of the gel.
- Record your results.
- Take a picture and label each well of the gel with the name of the student who's DNA was used, if the DNA was digested or not, and the genotype shown by the gel.
- Upload the picture to Canvas.
/Volumes/group/*SEP Files/*Uber SEP = Index/*Curriculum Updates/2017 Curriculum Update/PCR/TechGuide PCR 171010.docx 10/3/17Page 6 of 6
Proudly powered by Weebly